overview
Wisam
2022-02-12
phyloseq object
OTU table should be prepared, metadata and taxonomic table details
Example: phyloseq
The diet swap data set represents a study with African and African American groups undergoing a two-week diet swap. For details, see https://www.nature.com/articles/ncomms7342.
phyloseq-class experiment-level object
otu_table() OTU Table: [ 130 taxa and 222 samples ]
sample_data() Sample Data: [ 222 samples by 8 sample variables ]
tax_table() Taxonomy Table: [ 130 taxa by 3 taxonomic ranks ]Looking into the data
Number of taxonomic groups: 130
Number of samples: 222
Number of metadata fields: 8:
Metadata variables are: subject, sex, nationality, group, sample, timepoint, timepoint.within.group, bmi_group
Unique taxa at Phylum:
| phyloseq::get_taxa_unique(phy, “Phylum”) |
|---|
| Actinobacteria |
| Firmicutes |
| Proteobacteria |
| Verrucomicrobia |
| Bacteroidetes |
| Spirochaetes |
| Fusobacteria |
| Cyanobacteria |
Unique taxa at Genus:
| phyloseq::get_taxa_unique(phy, “Genus”) |
|---|
| Actinomycetaceae |
| Aerococcus |
| Aeromonas |
| Akkermansia |
| Alcaligenes faecalis et rel. |
| Allistipes et rel. |
| Anaerobiospirillum |
| Anaerofustis |
| Anaerostipes caccae et rel. |
| Anaerotruncus colihominis et rel. |
| Anaerovorax odorimutans et rel. |
| Aneurinibacillus |
| Aquabacterium |
| Asteroleplasma et rel. |
| Atopobium |
| Bacillus |
| Bacteroides fragilis et rel. |
| Bacteroides intestinalis et rel. |
| Bacteroides ovatus et rel. |
| Bacteroides plebeius et rel. |
| Bacteroides splachnicus et rel. |
| Bacteroides stercoris et rel. |
| Bacteroides uniformis et rel. |
| Bacteroides vulgatus et rel. |
| Bifidobacterium |
| Bilophila et rel. |
| Brachyspira |
| Bryantella formatexigens et rel. |
| Bulleidia moorei et rel. |
| Burkholderia |
| Butyrivibrio crossotus et rel. |
| Campylobacter |
| Catenibacterium mitsuokai et rel. |
| Clostridium (sensu stricto) |
| Clostridium cellulosi et rel. |
| Clostridium colinum et rel. |
| Clostridium difficile et rel. |
| Clostridium felsineum et rel. |
| Clostridium leptum et rel. |
| Clostridium nexile et rel. |
| Clostridium orbiscindens et rel. |
| Clostridium ramosum et rel. |
| Clostridium sphenoides et rel. |
| Clostridium stercorarium et rel. |
| Clostridium symbiosum et rel. |
| Clostridium thermocellum et rel. |
| Collinsella |
| Coprobacillus catenaformis et rel. |
| Coprococcus eutactus et rel. |
| Corynebacterium |
| Desulfovibrio et rel. |
| Dialister |
| Dorea formicigenerans et rel. |
| Eggerthella lenta et rel. |
| Enterobacter aerogenes et rel. |
| Enterococcus |
| Escherichia coli et rel. |
| Eubacterium biforme et rel. |
| Eubacterium cylindroides et rel. |
| Eubacterium hallii et rel. |
| Eubacterium limosum et rel. |
| Eubacterium rectale et rel. |
| Eubacterium siraeum et rel. |
| Eubacterium ventriosum et rel. |
| Faecalibacterium prausnitzii et rel. |
| Fusobacteria |
| Gemella |
| Granulicatella |
| Haemophilus |
| Helicobacter |
| Klebisiella pneumoniae et rel. |
| Lachnobacillus bovis et rel. |
| Lachnospira pectinoschiza et rel. |
| Lactobacillus catenaformis et rel. |
| Lactobacillus gasseri et rel. |
| Lactobacillus plantarum et rel. |
| Lactobacillus salivarius et rel. |
| Lactococcus |
| Leminorella |
| Megamonas hypermegale et rel. |
| Megasphaera elsdenii et rel. |
| Methylobacterium |
| Micrococcaceae |
| Mitsuokella multiacida et rel. |
| Moraxellaceae |
| Novosphingobium |
| Oceanospirillum |
| Oscillospira guillermondii et rel. |
| Outgrouping clostridium cluster XIVa |
| Oxalobacter formigenes et rel. |
| Papillibacter cinnamivorans et rel. |
| Parabacteroides distasonis et rel. |
| Peptococcus niger et rel. |
| Peptostreptococcus anaerobius et rel. |
| Peptostreptococcus micros et rel. |
| Phascolarctobacterium faecium et rel. |
| Prevotella melaninogenica et rel. |
| Prevotella oralis et rel. |
| Prevotella ruminicola et rel. |
| Prevotella tannerae et rel. |
| Propionibacterium |
| Proteus et rel. |
| Pseudomonas |
| Roseburia intestinalis et rel. |
| Ruminococcus bromii et rel. |
| Ruminococcus callidus et rel. |
| Ruminococcus gnavus et rel. |
| Ruminococcus lactaris et rel. |
| Ruminococcus obeum et rel. |
| Serratia |
| Sporobacter termitidis et rel. |
| Staphylococcus |
| Streptococcus bovis et rel. |
| Streptococcus intermedius et rel. |
| Streptococcus mitis et rel. |
| Subdoligranulum variable at rel. |
| Sutterella wadsworthia et rel. |
| Tannerella et rel. |
| Uncultured Bacteroidetes |
| Uncultured Chroococcales |
| Uncultured Clostridiales I |
| Uncultured Clostridiales II |
| Uncultured Mollicutes |
| Uncultured Selenomonadaceae |
| Veillonella |
| Vibrio |
| Weissella et rel. |
| Wissella et rel. |
| Xanthomonadaceae |
| Yersinia et rel. |
Useful plots
The taxa ranks in the previous phyloseq dataset are: Phylum, Family, Genus
Important to remember taxa abundances are compositional that are need appropriate transform to be informative, here I will use the relative abundances via compositional transform.
Abundance at Phylum level
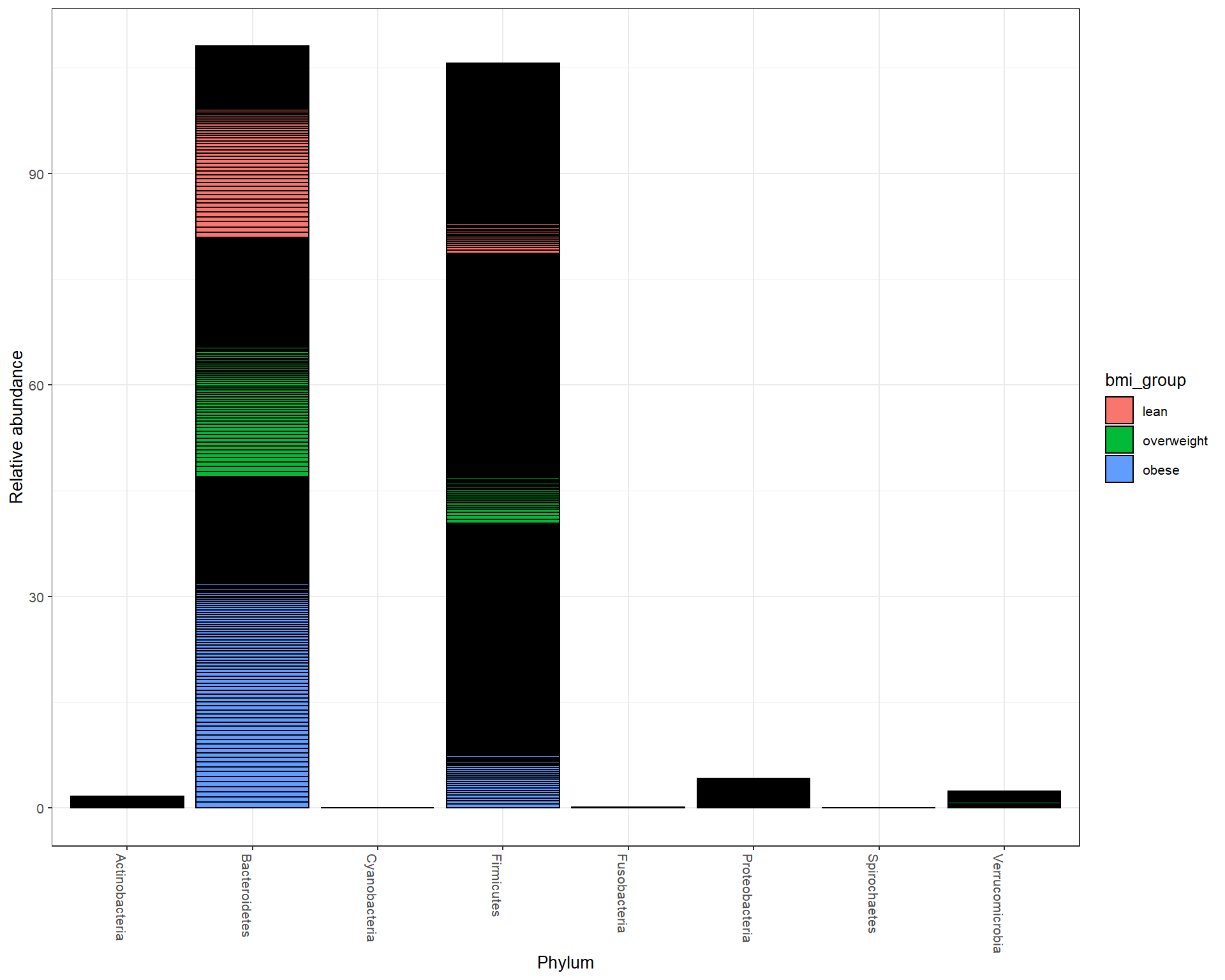
Abundance at Family level
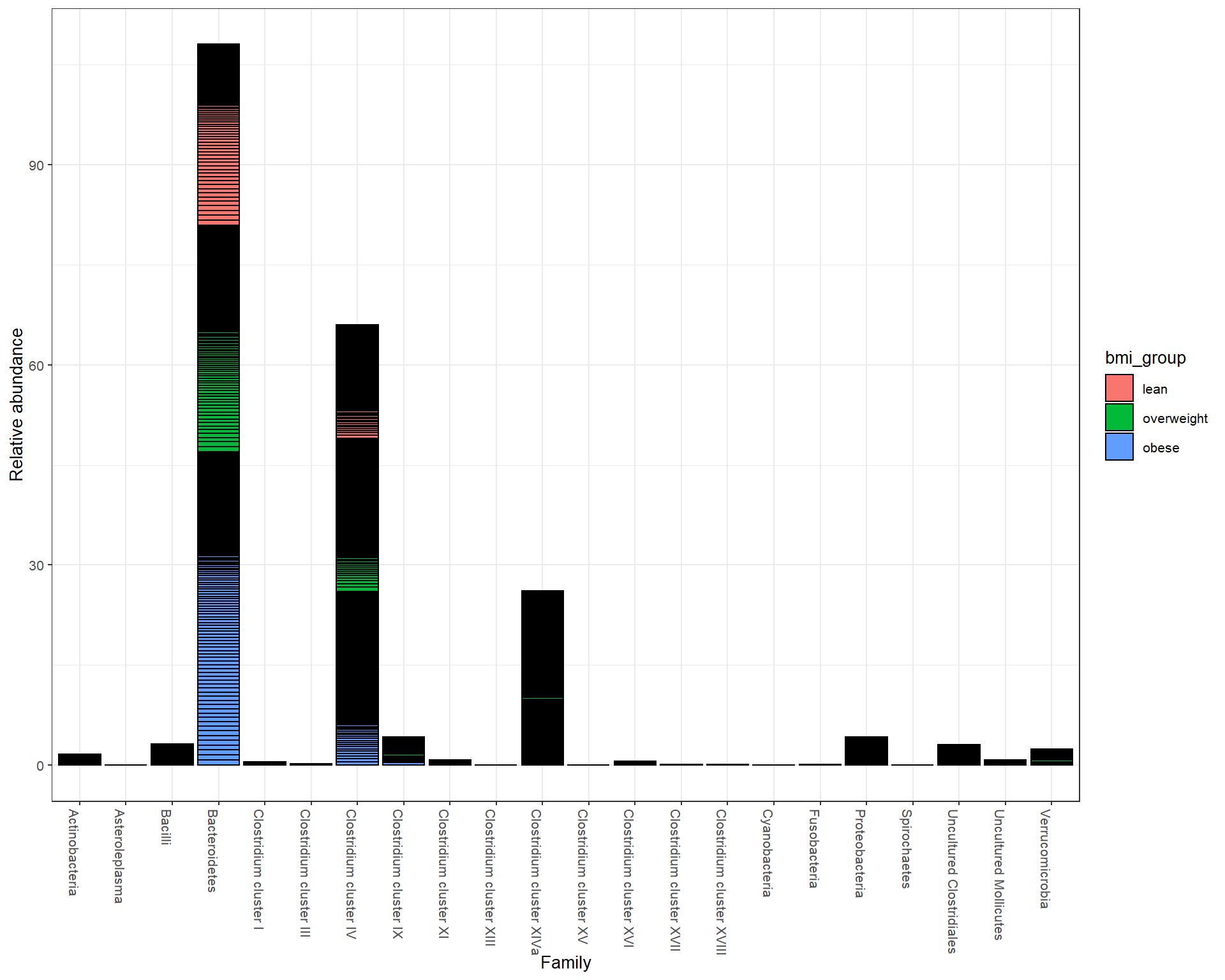
Abundance at Genus level
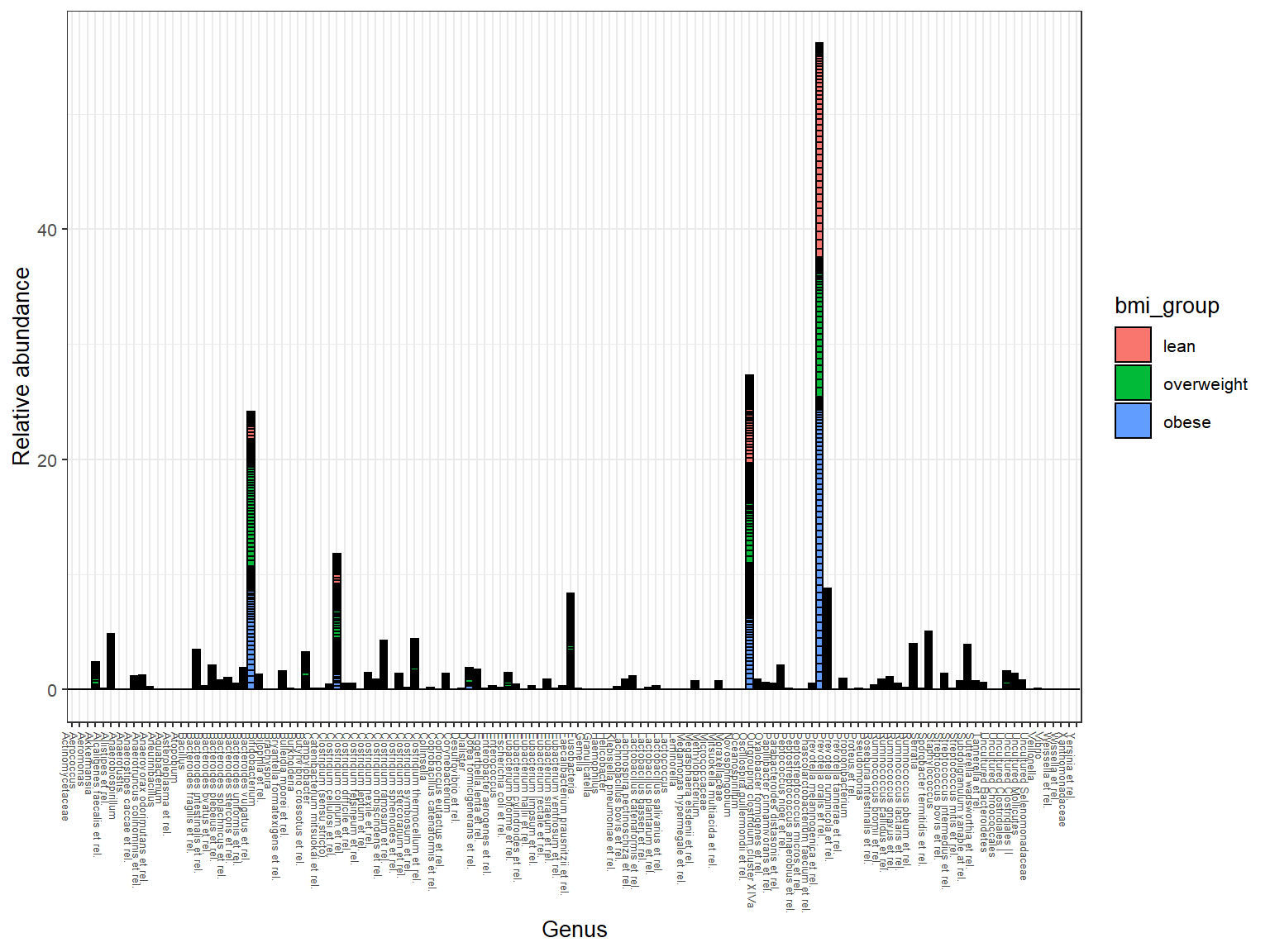
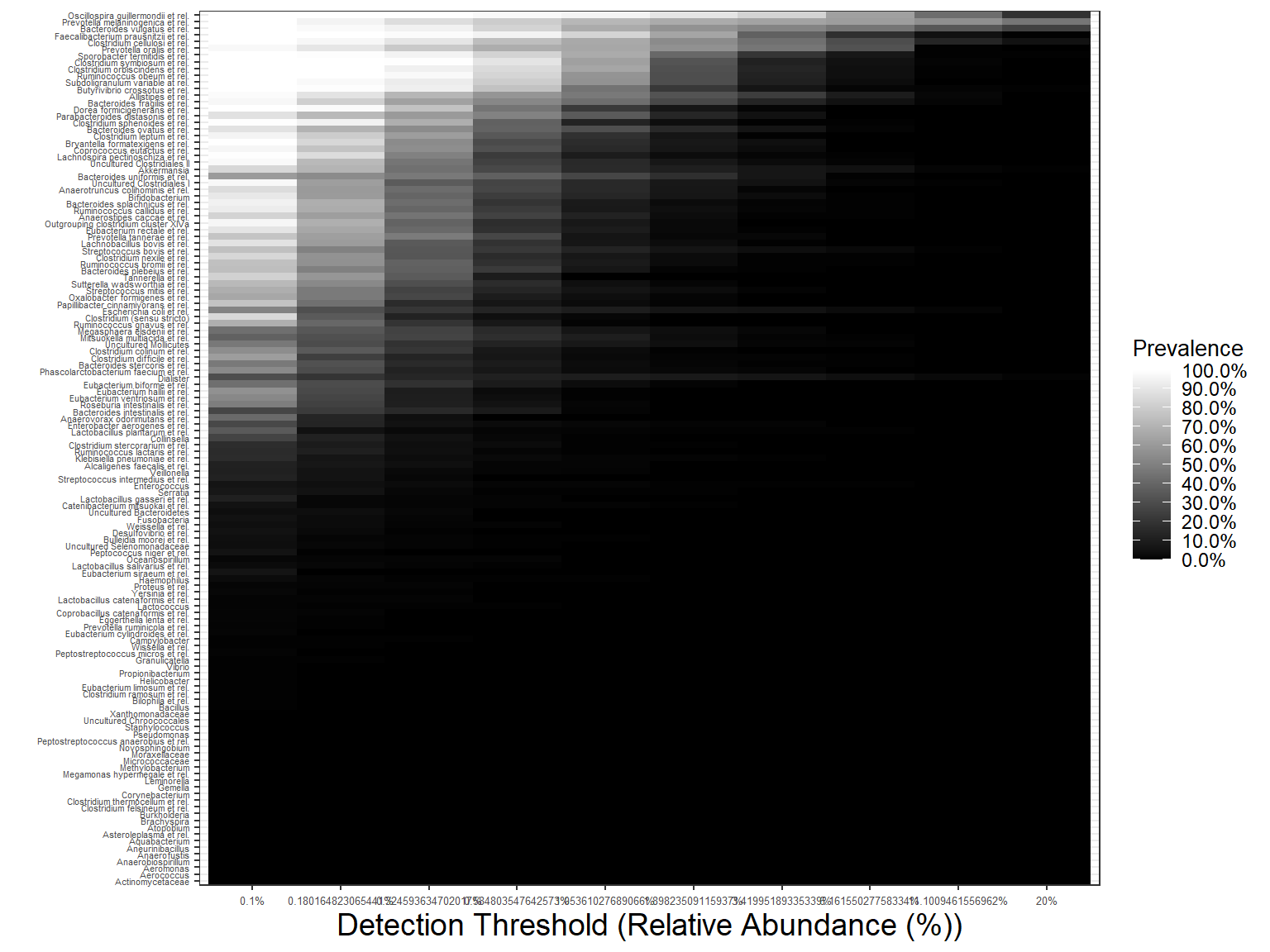
Heatmaps for the core taxa with certain prevalences & detections
Core heatmaps
Acknowledgement to "Leo Lahti, Sudarshan Shetty et al. (2017). Tools for microbiome analysis in R. Version . URL: http://microbiome.github.com/microbiome