Installing ALDEx2
ALDEx2 is a bioconductor package that could be installed Via bioconductor
library(BiocManager)
#BiocManager::install('ALDEx2')ALDEx2 tutorials
ALDEx2 uses non-parametric statistical modeling
Useful book
Working with ALDEx2
Dataset dietswap
- Phyloseq dataset From microbiome package dietswap
library(microbiome)
data("dietswap")
dietswapphyloseq-class experiment-level object
otu_table() OTU Table: [ 130 taxa and 222 samples ]
sample_data() Sample Data: [ 222 samples by 8 sample variables ]
tax_table() Taxonomy Table: [ 130 taxa by 3 taxonomic ranks ]check the number of genera 130
check the number of samples 222
The two groups of interest in our Aldex2 analysis are:
AAM, it means African American
AFR, it means African
The whole variables of dietswap dataset are: StCode, StVisit, InterventionGroup, MGDMOGTT1OR2Fi, MGDM1Fi, MGDM2FiNEW, MPrSerumMMP8, MPrSerumphIGFBP1, MPrSerumIGFBP1, MCRP, MPrVagMMP8, MAge1, MprepBMI, MBMIOverwOrObese, MLivingStatus, MUniEdu, MEdu, MPA, MMETINDX, MprepSmoke, MSmoke, MEthnicity, MEkJ, MEkcal, MProg, MHHg, MFatg, MFiballg, ProEPercent, MHHEPercent, MFatEPercent, Mparity, MPRW, MPrAbVag, MPrAbVag4weeks, MPrVaginaTreatment, MPrVaginaTreatmentTotalNo, MPrVaginaTreatment4weeks, MPRAb, MPRAbNo, MPRAb4weeks, MPrevProb1, MPrevProb1withintwoweeks, MPrevProb1beforetwoweeks, MProbUse, MBleedElevBBorAnemia1, allGroups, group
Main tests
t-test
T-test between groups where wi.eBH<= 0.05 and we.eBH <= 0.05. The first tens only.
| wi.eBH | we.eBH | |
|---|---|---|
| Allistipes et rel. | 0.0000000 | 0.0000000 |
| Anaerostipes caccae et rel. | 0.0004071 | 0.0021022 |
| Bacteroides fragilis et rel. | 0.0000252 | 0.0000110 |
| Bacteroides intestinalis et rel. | 0.0000009 | 0.0000026 |
| Bacteroides ovatus et rel. | 0.0000000 | 0.0000000 |
| Bacteroides plebeius et rel. | 0.0000000 | 0.0000000 |
| Bacteroides splachnicus et rel. | 0.0001776 | 0.0059688 |
| Bacteroides stercoris et rel. | 0.0000004 | 0.0000482 |
| Bacteroides uniformis et rel. | 0.0000031 | 0.0000006 |
| Bacteroides vulgatus et rel. | 0.0000000 | 0.0000000 |
Kruskal-Wallis test
Kruskal-Wallis test between groups where kw.eBH <=0.05. The first tens only.
| kw.eBH | kw.ep | |
|---|---|---|
| Allistipes et rel. | 0.0000000 | 0.00e+00 |
| Anaerostipes caccae et rel. | 0.0004071 | 7.16e-05 |
| Bacteroides fragilis et rel. | 0.0000252 | 3.00e-06 |
| Bacteroides intestinalis et rel. | 0.0000009 | 1.00e-07 |
| Bacteroides ovatus et rel. | 0.0000000 | 0.00e+00 |
| Bacteroides plebeius et rel. | 0.0000000 | 0.00e+00 |
| Bacteroides splachnicus et rel. | 0.0001776 | 2.93e-05 |
| Bacteroides stercoris et rel. | 0.0000004 | 0.00e+00 |
| Bacteroides uniformis et rel. | 0.0000031 | 3.00e-07 |
| Bacteroides vulgatus et rel. | 0.0000000 | 0.00e+00 |
Estimate Effect Size
Estimate Effect Size between groups where the effect > 0.1 or the effect < -0.1, the first tens only.
| rab.all | rab.win.AAM | rab.win.AFR | diff.btw | diff.win | effect | overlap | |
|---|---|---|---|---|---|---|---|
| Akkermansia | 2.4333721 | 2.3264607 | 2.5920414 | 0.2718289 | 2.331340 | 0.1003826 | 0.4468000 |
| Alcaligenes faecalis et rel. | -1.2698393 | -1.3608975 | -1.1478797 | 0.2714610 | 2.366439 | 0.1012274 | 0.4502000 |
| Allistipes et rel. | 3.8782135 | 3.1801389 | 5.1119393 | 1.6625243 | 2.404121 | 0.6552559 | 0.2383523 |
| Anaerostipes caccae et rel. | 2.3541216 | 1.8667819 | 2.8020553 | 0.7463802 | 1.829484 | 0.3672804 | 0.3371326 |
| Anaerotruncus colihominis et rel. | 2.1989229 | 2.4109583 | 1.9022760 | -0.2617569 | 2.225332 | -0.1136506 | 0.4465107 |
| Anaerovorax odorimutans et rel. | 0.7969434 | 0.9100502 | 0.6716430 | -0.2617360 | 1.194559 | -0.2045655 | 0.4057189 |
| Bacteroides fragilis et rel. | 3.6851499 | 2.7277086 | 4.4171758 | 1.1424114 | 2.570196 | 0.4303816 | 0.3289342 |
| Bacteroides intestinalis et rel. | -0.6130119 | -1.2377048 | 0.3646694 | 1.6472656 | 3.126962 | 0.5076885 | 0.2738000 |
| Bacteroides ovatus et rel. | 2.8630332 | 2.0412264 | 3.9044801 | 1.5391897 | 2.213514 | 0.6573121 | 0.2384000 |
| Bacteroides plebeius et rel. | 1.9390361 | 1.2653126 | 2.5952103 | 1.2614453 | 1.784423 | 0.6575322 | 0.2352000 |
Summary
The first 15 only.
| diff.btw | diff.win | effect | overlap | we.ep | wi.ep | we.eBH | wi.eBH | |
|---|---|---|---|---|---|---|---|---|
| Allistipes et rel. | 1.6625243 | 2.404121 | 0.6552559 | 0.2383523 | 0.0000000 | 0.0000000 | 0.0000000 | 0.0000000 |
| Anaerostipes caccae et rel. | 0.7463802 | 1.829484 | 0.3672804 | 0.3371326 | 0.0004109 | 0.0000716 | 0.0021022 | 0.0004071 |
| Anaerovorax odorimutans et rel. | -0.2617360 | 1.194559 | -0.2045655 | 0.4057189 | 0.0272605 | 0.0473909 | 0.0620924 | 0.1014251 |
| Bacteroides fragilis et rel. | 1.1424114 | 2.570196 | 0.4303816 | 0.3289342 | 0.0000012 | 0.0000030 | 0.0000110 | 0.0000252 |
| Bacteroides intestinalis et rel. | 1.6472656 | 3.126962 | 0.5076885 | 0.2738000 | 0.0000003 | 0.0000001 | 0.0000026 | 0.0000009 |
| Bacteroides ovatus et rel. | 1.5391897 | 2.213514 | 0.6573121 | 0.2384000 | 0.0000000 | 0.0000000 | 0.0000000 | 0.0000000 |
| Bacteroides plebeius et rel. | 1.2614453 | 1.784423 | 0.6575322 | 0.2352000 | 0.0000000 | 0.0000000 | 0.0000000 | 0.0000000 |
| Bacteroides splachnicus et rel. | 0.5558292 | 1.407581 | 0.3456709 | 0.3274000 | 0.0013684 | 0.0000293 | 0.0059688 | 0.0001776 |
| Bacteroides stercoris et rel. | 1.0565405 | 1.807949 | 0.5030065 | 0.2749450 | 0.0000068 | 0.0000000 | 0.0000482 | 0.0000004 |
| Bacteroides uniformis et rel. | 1.8479531 | 3.673068 | 0.4947658 | 0.2981404 | 0.0000000 | 0.0000003 | 0.0000006 | 0.0000031 |
| Bacteroides vulgatus et rel. | 2.3028431 | 3.278722 | 0.6886983 | 0.2447511 | 0.0000000 | 0.0000000 | 0.0000000 | 0.0000000 |
| Bifidobacterium | 0.5896979 | 1.978390 | 0.2738965 | 0.3558000 | 0.0041537 | 0.0005298 | 0.0154198 | 0.0025087 |
| Bryantella formatexigens et rel. | 0.3458719 | 1.650042 | 0.2012445 | 0.4017197 | 0.0139609 | 0.0108863 | 0.0428020 | 0.0355833 |
| Bulleidia moorei et rel. | -0.6576048 | 1.708306 | -0.3391389 | 0.3370000 | 0.0002358 | 0.0007945 | 0.0009883 | 0.0028710 |
| Catenibacterium mitsuokai et rel. | -1.4408704 | 3.525224 | -0.3484797 | 0.3212000 | 0.0003128 | 0.0000697 | 0.0014030 | 0.0003602 |
ALDEx2 Wrapper
The first 15 only.
| rab.all | rab.win.AAM | rab.win.AFR | diff.btw | diff.win | effect | overlap | we.ep | we.eBH | wi.ep | wi.eBH | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Actinomycetaceae | -5.4097921 | -5.416925 | -5.4025698 | 0.0460888 | 4.220298 | 0.0087526 | 0.4957009 | 0.5372296 | 0.6557136 | 0.5316953 | 0.6508055 |
| Aeromonas | -5.6117266 | -5.636775 | -5.5801023 | 0.0404607 | 4.290807 | 0.0083972 | 0.4955009 | 0.4886397 | 0.6195258 | 0.4908196 | 0.6220262 |
| Akkermansia | 1.6706992 | 1.544770 | 1.8222962 | 0.2904666 | 2.412044 | 0.1083721 | 0.4428000 | 0.1877045 | 0.3525808 | 0.1542397 | 0.3072821 |
| Alcaligenes faecalis et rel. | -2.0095630 | -2.130209 | -1.8519295 | 0.3554576 | 2.492967 | 0.1294209 | 0.4370000 | 0.1444239 | 0.2619220 | 0.2165351 | 0.3476054 |
| Allistipes et rel. | 3.1790350 | 2.469320 | 4.4031057 | 1.6721978 | 2.517624 | 0.6421007 | 0.2456000 | 0.0000000 | 0.0000000 | 0.0000000 | 0.0000000 |
| Anaerobiospirillum | -5.5786364 | -5.598406 | -5.5582017 | 0.1019949 | 4.195919 | 0.0215416 | 0.4884000 | 0.5111306 | 0.6354885 | 0.4950349 | 0.6224329 |
| Anaerofustis | -5.5107962 | -5.633154 | -5.3493605 | 0.2908242 | 4.219219 | 0.0568741 | 0.4698000 | 0.4442366 | 0.5705238 | 0.4198762 | 0.5558390 |
| Anaerostipes caccae et rel. | 1.5842199 | 1.172157 | 2.0647986 | 0.7505529 | 1.754072 | 0.3929324 | 0.3264000 | 0.0001496 | 0.0008357 | 0.0000346 | 0.0002257 |
| Anaerotruncus colihominis et rel. | 1.4724702 | 1.643813 | 1.2286622 | -0.2234073 | 2.213632 | -0.0906825 | 0.4548000 | 0.4076705 | 0.5825280 | 0.2644596 | 0.4406690 |
| Anaerovorax odorimutans et rel. | 0.0534992 | 0.173551 | -0.0880767 | -0.2507698 | 1.207598 | -0.1923404 | 0.4093181 | 0.0360819 | 0.0767754 | 0.0490105 | 0.1021640 |
| Aquabacterium | -5.2878133 | -5.192724 | -5.3970588 | -0.2566430 | 4.303852 | -0.0516166 | 0.4707059 | 0.4371709 | 0.5713546 | 0.4429046 | 0.5751277 |
| Atopobium | -5.2638824 | -5.201397 | -5.3499520 | -0.1134649 | 4.132625 | -0.0238986 | 0.4860000 | 0.4897667 | 0.6157344 | 0.4960600 | 0.6210132 |
| Bacillus | -3.2284247 | -3.248051 | -3.2063139 | 0.0257078 | 2.119265 | 0.0112742 | 0.4945011 | 0.5640247 | 0.6868960 | 0.5521835 | 0.6758519 |
| Bacteroides fragilis et rel. | 2.9086180 | 1.986017 | 3.6797470 | 1.1709792 | 2.635873 | 0.4268490 | 0.3242000 | 0.0000029 | 0.0000270 | 0.0000070 | 0.0000605 |
| Bacteroides intestinalis et rel. | -1.3242737 | -1.964276 | -0.3196531 | 1.6777492 | 3.196426 | 0.5060486 | 0.2837433 | 0.0000017 | 0.0000120 | 0.0000002 | 0.0000020 |
Plots
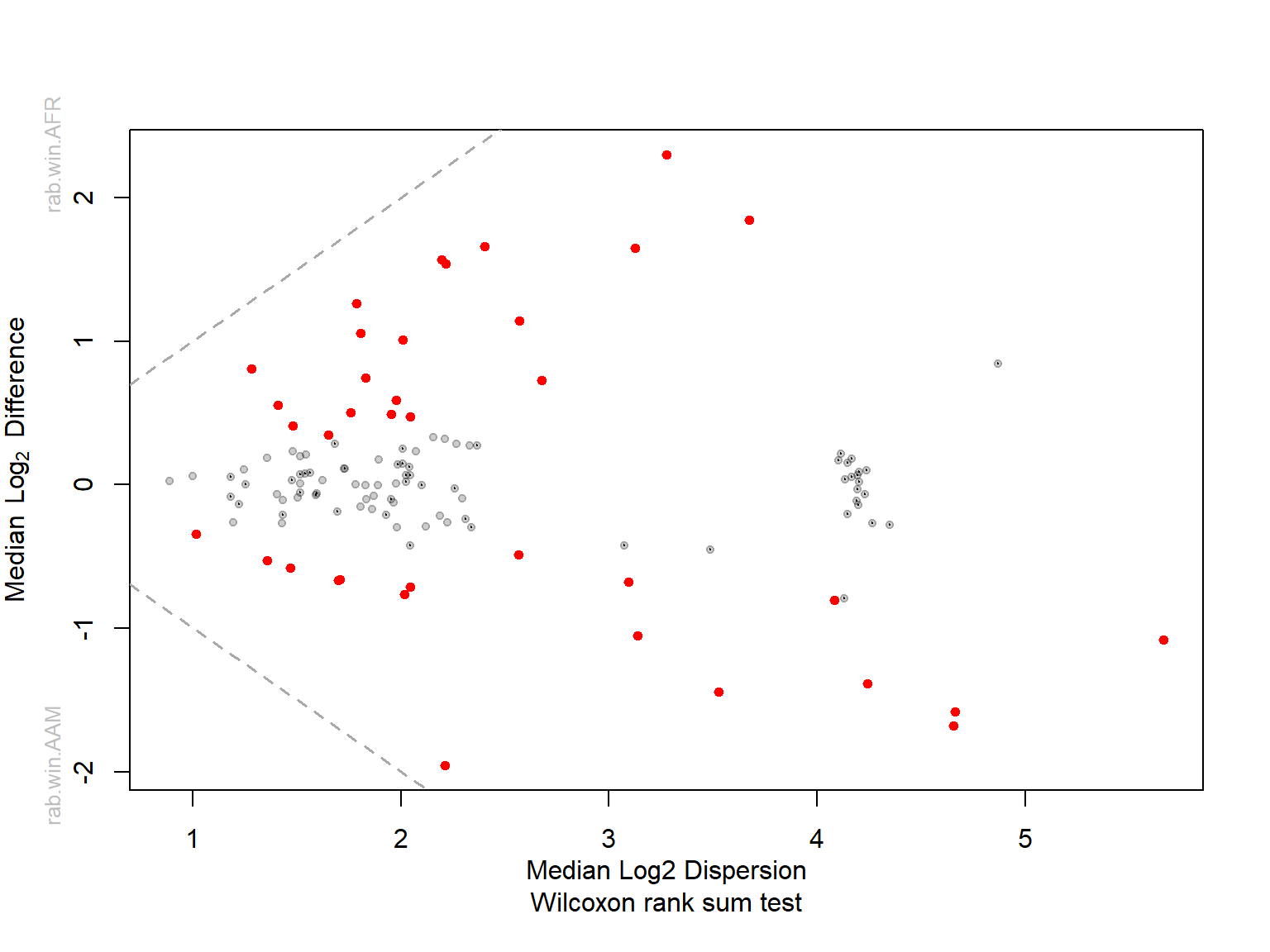
Bland-Altman plot
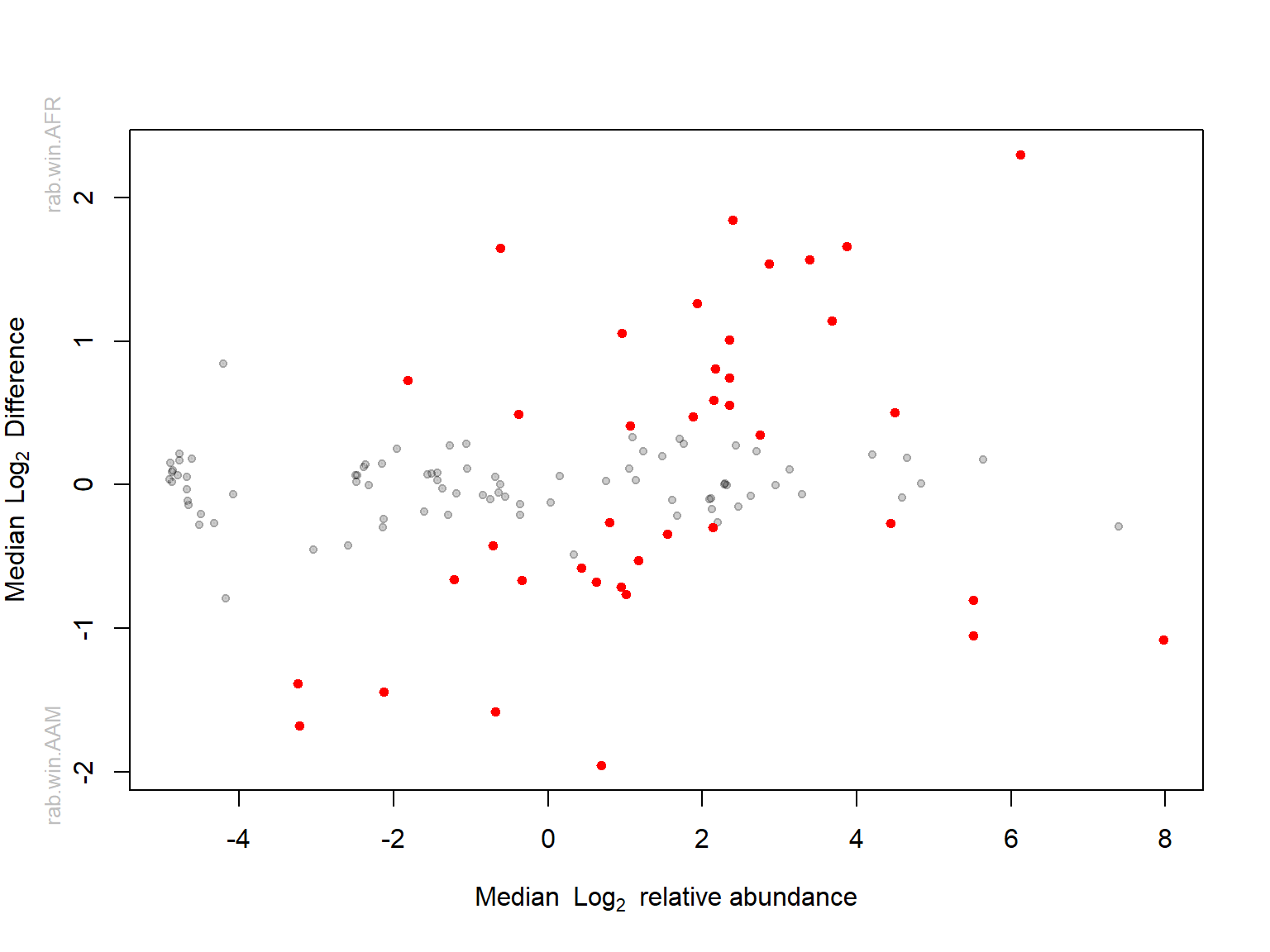
Red taxa are the significant ones with adjusted p-values cutoff 0.05 or less.
Grey taxa are the rare ones
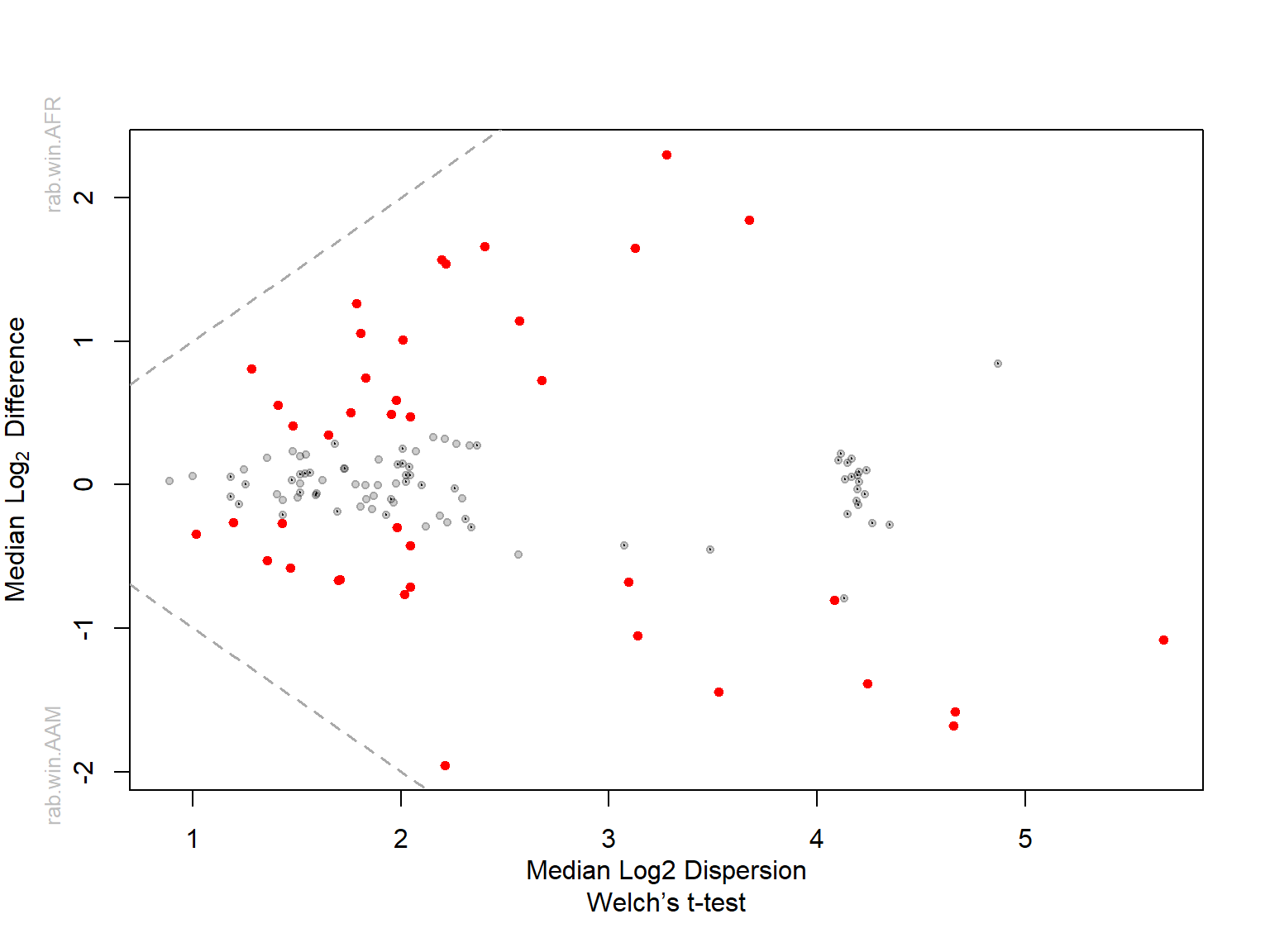
Median plots
Plot median between-group difference Versus median within-group difference
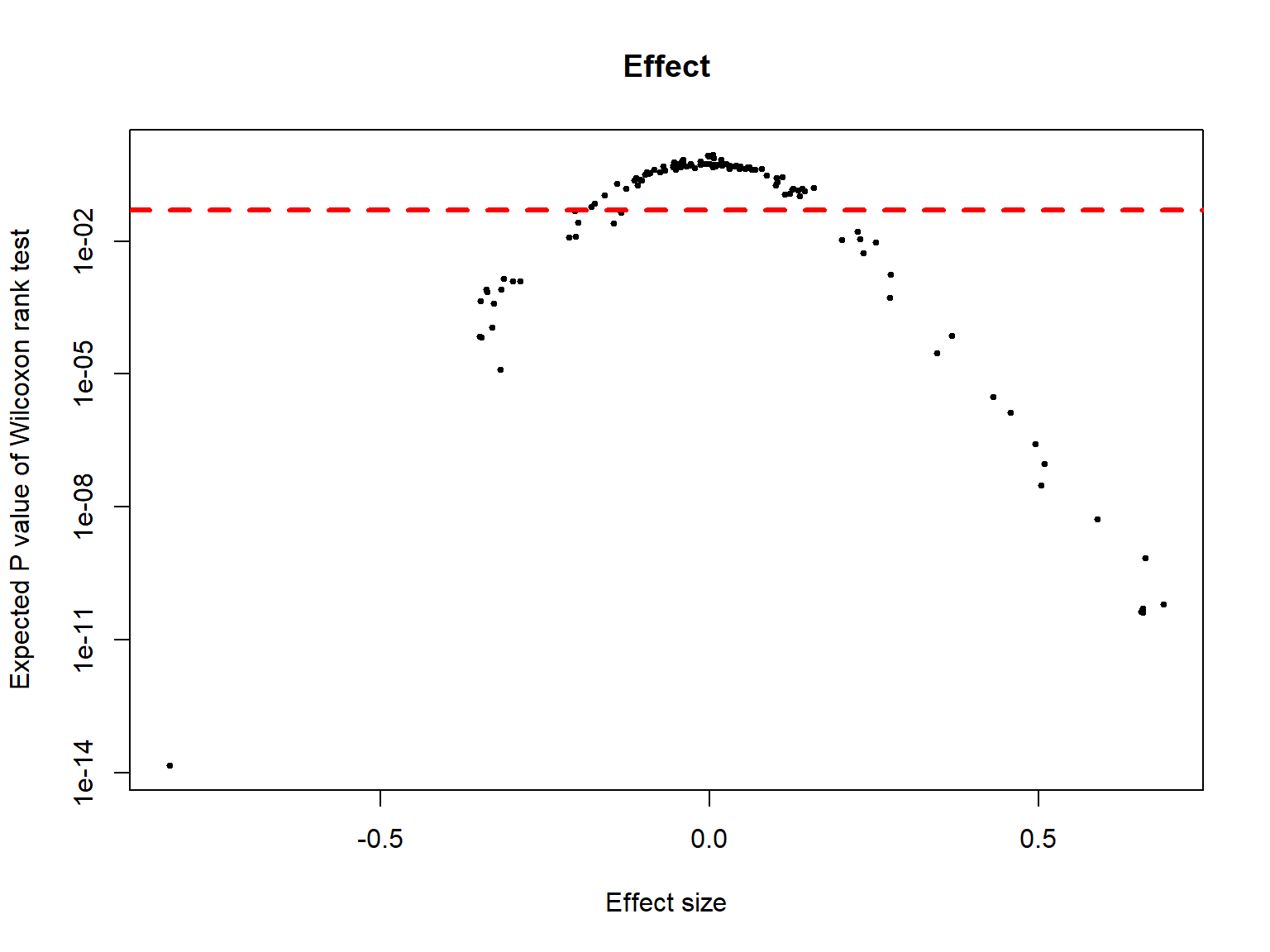
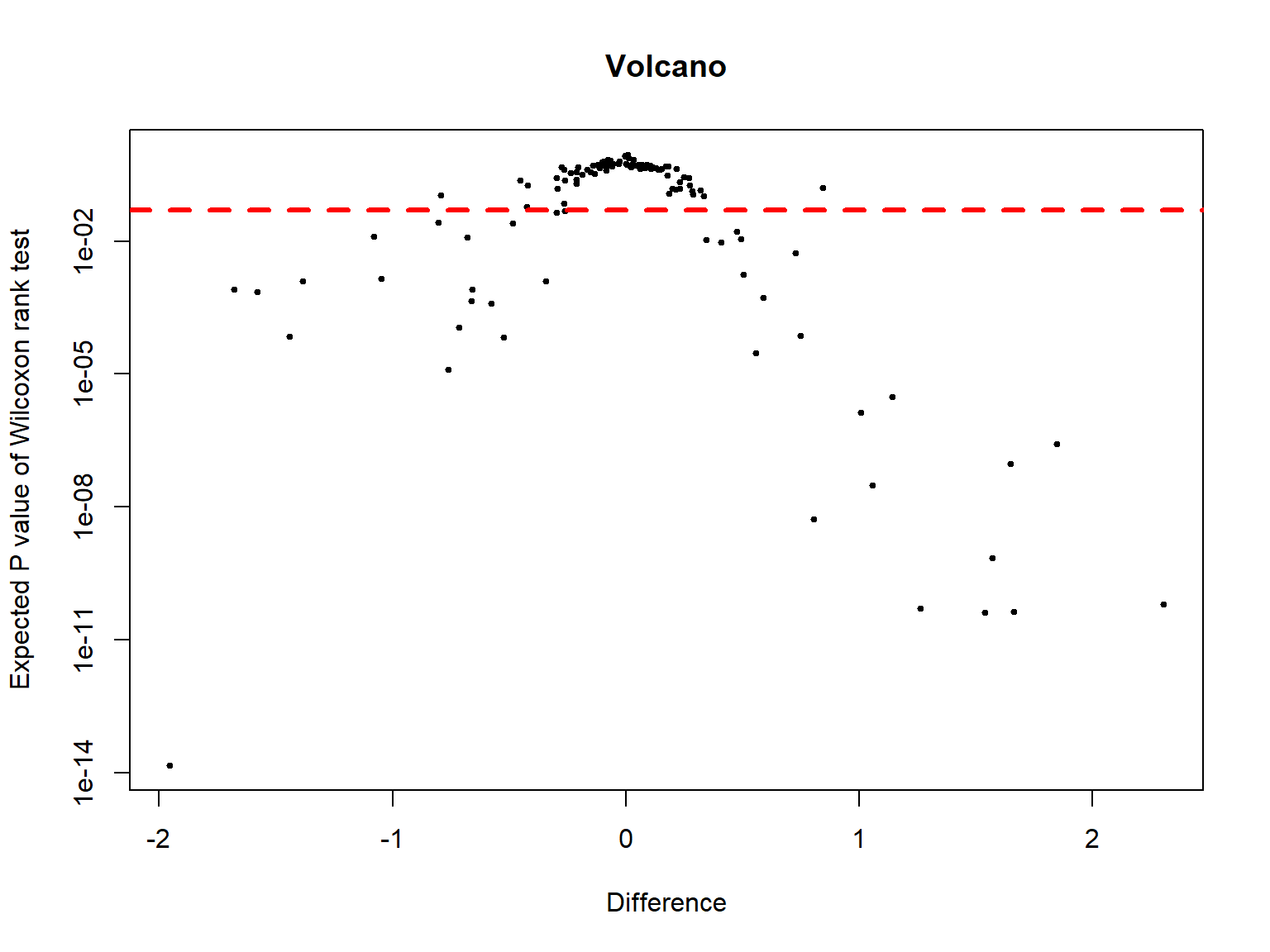
Effect and Volcano plots